Validating Sam's data with GoodTables
Introduction
As a human microbiome researcher, I don't often get a chance to play with non-sequencing data. I'm excited to try my hand at handling and validating other types of data. In this post, I'll be validating data with GoodTables from fellow westcoaster, Sam Wilairat. She's already written about her experience with GoodTables here, and I'll be trying to recreate what she did.
Sam's Data
Sam gave me a CSV file that is a dataset with Health Science Open Education Resources (OER). She created this dataset so that students, educators, clinicians, and the general public can more easily find and utilize free learning tools. The file data includes not only the title and author of the resource, but also the url where it's freely available.
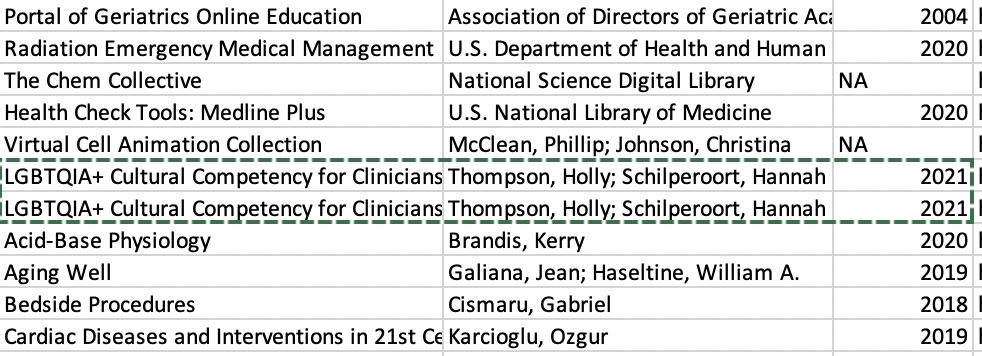
To test the capabilities of GoodTables validation, Sam has duplicated one of the rows of data (LGBTQIA+ Cultural Competency for Clinicians), shown below:
GoodTables time
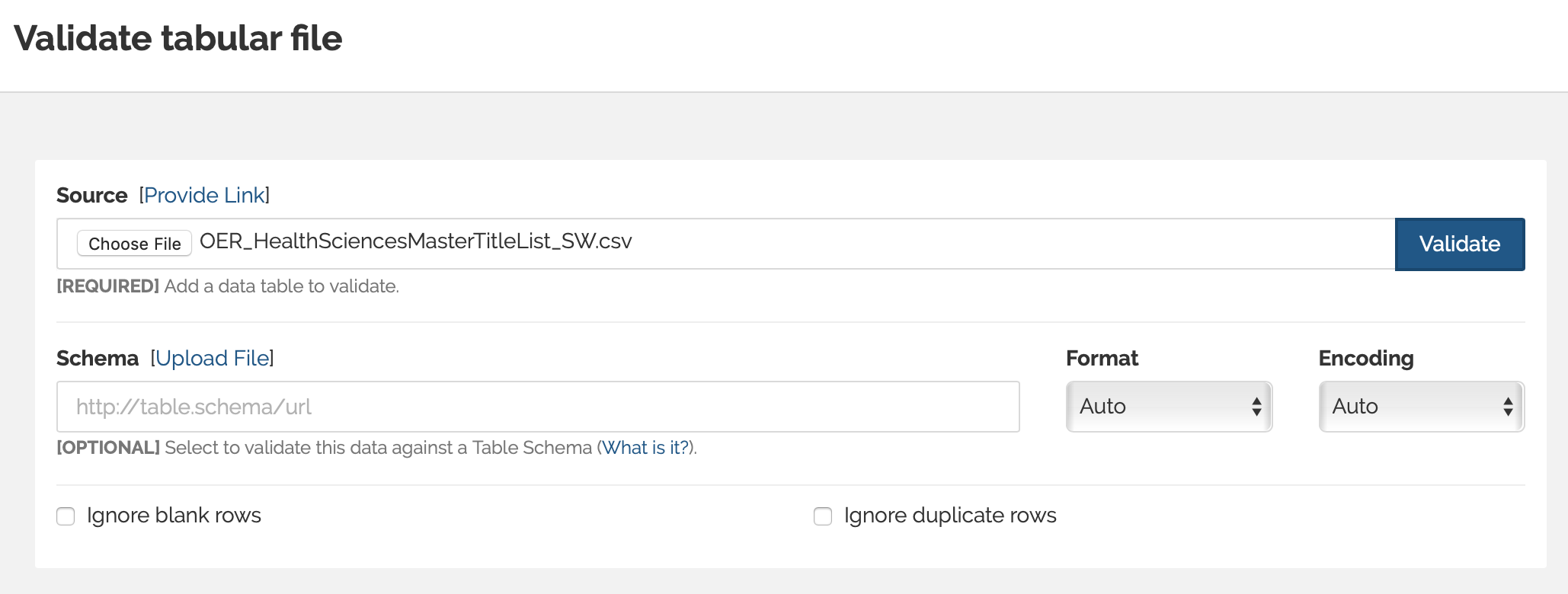
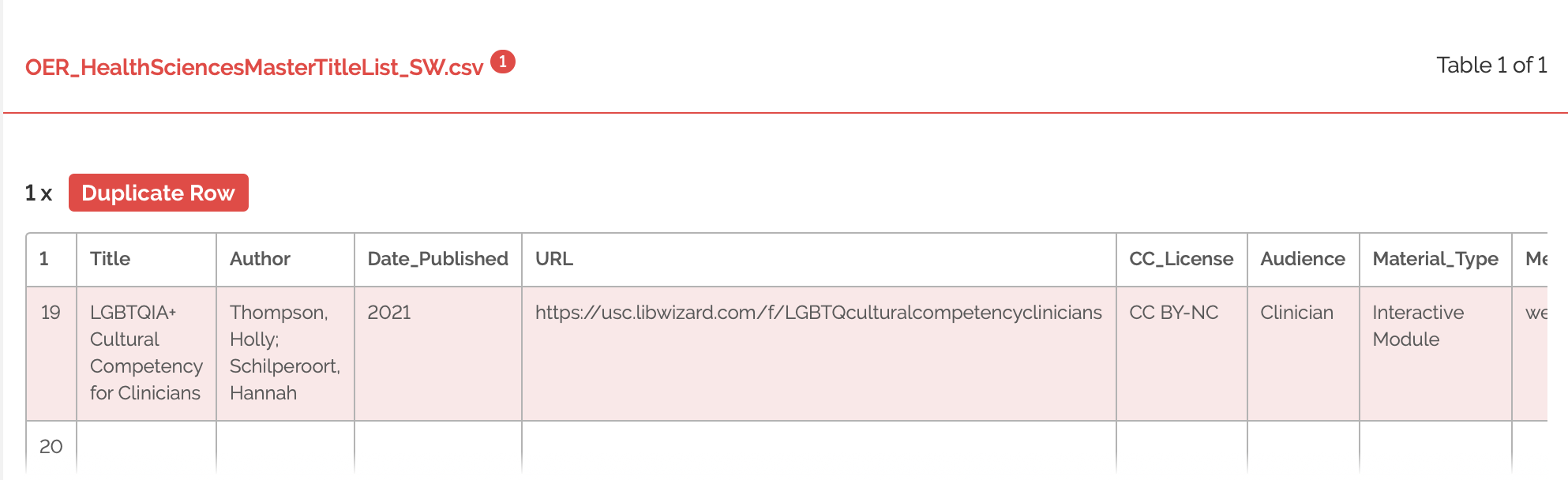
Sam used both the browser tool and command line GoodTables interface. Starting with the browser tool, I uploaded the csv file as the source file. As expected, the GoodTables browser tool detects the duplicated rows and throws an error.
Moving on to the command line tool for GoodTables. I had yet to play around with this tool, so this was a good excuse to finally learn the command line interface. After installing the GoodTables tool using
pip install goodtables
I followed the instructions from Sam's blog, and tested GoodTables on the OER dataset with the duplicated row using:
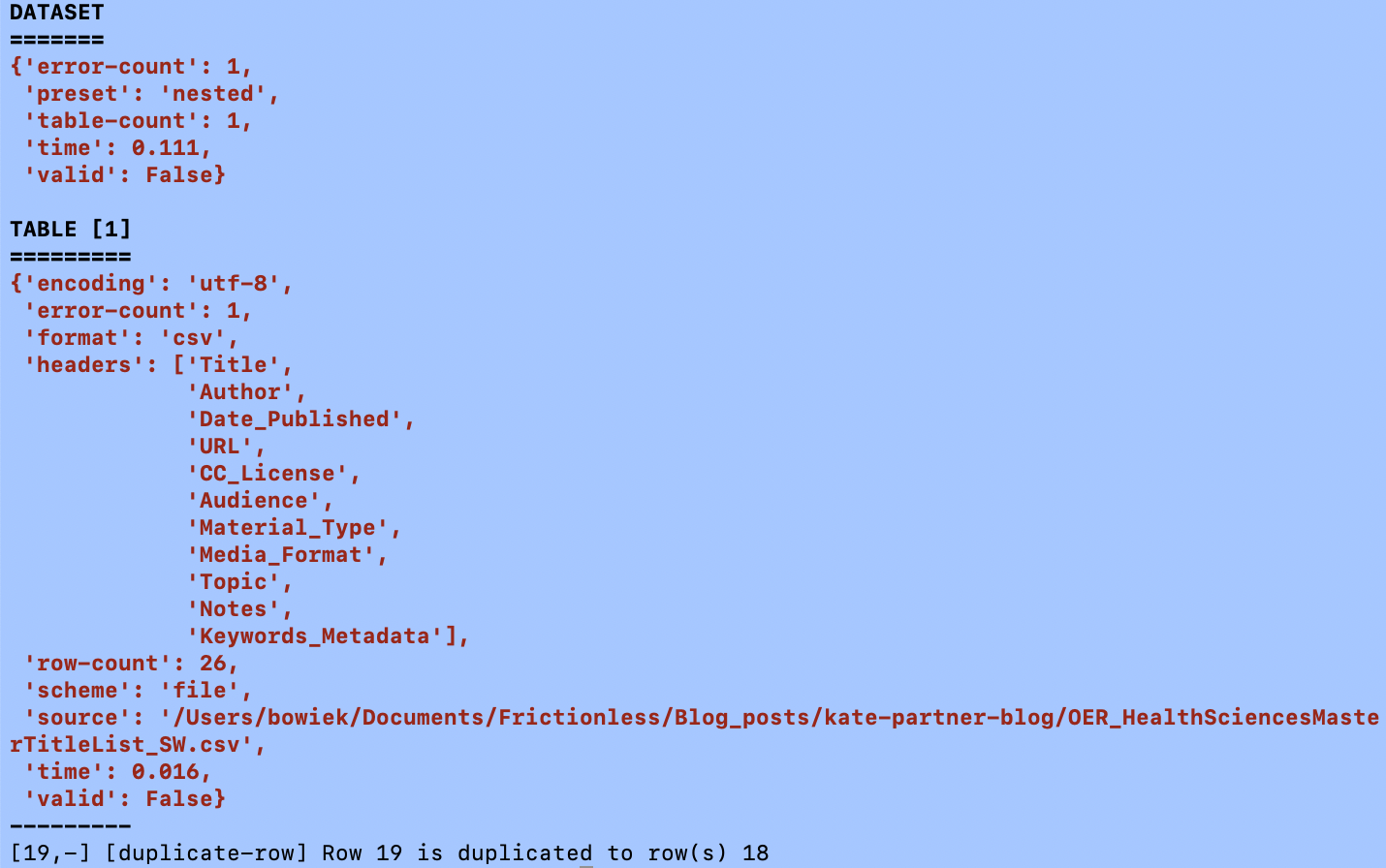
goodtables path/to/OER_HealthSciencesMasterTitleList_SW.csv
Again, as expected and described in Sam's earlier blog, the command line GoodTables interface also threw an error: